Phospho-p53 (Ser20) Antibody - #AF3073
製品説明
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
引用形式: Affinity Biosciences Cat# AF3073, RRID:AB_2834510.
折りたたみ/展開
Antigen NY-CO-13; BCC7; Cellular tumor antigen p53; FLJ92943; LFS1; Mutant tumor protein 53; p53; p53 tumor suppressor; P53_HUMAN; Phosphoprotein p53; Tp53; Transformation related protein 53; TRP53; Tumor protein 53; Tumor protein p53; Tumor suppressor p53;
免疫原
A synthesized peptide derived from human p53 around the phosphorylation site of Ser20.
Ubiquitous. Isoforms are expressed in a wide range of normal tissues but in a tissue-dependent manner. Isoform 2 is expressed in most normal tissues but is not detected in brain, lung, prostate, muscle, fetal brain, spinal cord and fetal liver. Isoform 3 is expressed in most normal tissues but is not detected in lung, spleen, testis, fetal brain, spinal cord and fetal liver. Isoform 7 is expressed in most normal tissues but is not detected in prostate, uterus, skeletal muscle and breast. Isoform 8 is detected only in colon, bone marrow, testis, fetal brain and intestine. Isoform 9 is expressed in most normal tissues but is not detected in brain, heart, lung, fetal liver, salivary gland, breast or intestine.
- P04637 P53_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MEEPQSDPSVEPPLSQETFSDLWKLLPENNVLSPLPSQAMDDLMLSPDDIEQWFTEDPGPDEAPRMPEAAPPVAPAPAAPTPAAPAPAPSWPLSSSVPSQKTYQGSYGFRLGFLHSGTAKSVTCTYSPALNKMFCQLAKTCPVQLWVDSTPPPGTRVRAMAIYKQSQHMTEVVRRCPHHERCSDSDGLAPPQHLIRVEGNLRVEYLDDRNTFRHSVVVPYEPPEVGSDCTTIHYNYMCNSSCMGGMNRRPILTIITLEDSSGNLLGRNSFEVRVCACPGRDRRTEEENLRKKGEPHHELPPGSTKRALPNNTSSSPQPKKKPLDGEYFTLQIRGRERFEMFRELNEALELKDAQAGKEPGGSRAHSSHLKSKKGQSTSRHKKLMFKTEGPDSD
種類予測
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
研究背景
Acts as a tumor suppressor in many tumor types; induces growth arrest or apoptosis depending on the physiological circumstances and cell type. Involved in cell cycle regulation as a trans-activator that acts to negatively regulate cell division by controlling a set of genes required for this process. One of the activated genes is an inhibitor of cyclin-dependent kinases. Apoptosis induction seems to be mediated either by stimulation of BAX and FAS antigen expression, or by repression of Bcl-2 expression. Its pro-apoptotic activity is activated via its interaction with PPP1R13B/ASPP1 or TP53BP2/ASPP2. However, this activity is inhibited when the interaction with PPP1R13B/ASPP1 or TP53BP2/ASPP2 is displaced by PPP1R13L/iASPP. In cooperation with mitochondrial PPIF is involved in activating oxidative stress-induced necrosis; the function is largely independent of transcription. Induces the transcription of long intergenic non-coding RNA p21 (lincRNA-p21) and lincRNA-Mkln1. LincRNA-p21 participates in TP53-dependent transcriptional repression leading to apoptosis and seems to have an effect on cell-cycle regulation. Implicated in Notch signaling cross-over. Prevents CDK7 kinase activity when associated to CAK complex in response to DNA damage, thus stopping cell cycle progression. Isoform 2 enhances the transactivation activity of isoform 1 from some but not all TP53-inducible promoters. Isoform 4 suppresses transactivation activity and impairs growth suppression mediated by isoform 1. Isoform 7 inhibits isoform 1-mediated apoptosis. Regulates the circadian clock by repressing CLOCK-ARNTL/BMAL1-mediated transcriptional activation of PER2.
Acetylated. Acetylation of Lys-382 by CREBBP enhances transcriptional activity. Deacetylation of Lys-382 by SIRT1 impairs its ability to induce proapoptotic program and modulate cell senescence. Deacetylation by SIRT2 impairs its ability to induce transcription activation in a AKT-dependent manner.
Phosphorylation on Ser residues mediates transcriptional activation. Phosphorylated by HIPK1 (By similarity). Phosphorylation at Ser-9 by HIPK4 increases repression activity on BIRC5 promoter. Phosphorylated on Thr-18 by VRK1. Phosphorylated on Ser-20 by CHEK2 in response to DNA damage, which prevents ubiquitination by MDM2. Phosphorylated on Ser-20 by PLK3 in response to reactive oxygen species (ROS), promoting p53/TP53-mediated apoptosis. Phosphorylated on Thr-55 by TAF1, which promotes MDM2-mediated degradation. Phosphorylated on Ser-33 by CDK7 in a CAK complex in response to DNA damage. Phosphorylated on Ser-46 by HIPK2 upon UV irradiation. Phosphorylation on Ser-46 is required for acetylation by CREBBP. Phosphorylated on Ser-392 following UV but not gamma irradiation. Phosphorylated on Ser-15 upon ultraviolet irradiation; which is enhanced by interaction with BANP. Phosphorylated by NUAK1 at Ser-15 and Ser-392; was initially thought to be mediated by STK11/LKB1 but it was later shown that it is indirect and that STK11/LKB1-dependent phosphorylation is probably mediated by downstream NUAK1. It is unclear whether AMP directly mediates phosphorylation at Ser-15. Phosphorylated on Thr-18 by isoform 1 and isoform 2 of VRK2. Phosphorylation on Thr-18 by isoform 2 of VRK2 results in a reduction in ubiquitination by MDM2 and an increase in acetylation by EP300. Stabilized by CDK5-mediated phosphorylation in response to genotoxic and oxidative stresses at Ser-15, Ser-33 and Ser-46, leading to accumulation of p53/TP53, particularly in the nucleus, thus inducing the transactivation of p53/TP53 target genes. Phosphorylated by DYRK2 at Ser-46 in response to genotoxic stress. Phosphorylated at Ser-315 and Ser-392 by CDK2 in response to DNA-damage. Phosphorylation at Ser-15 is required for interaction with DDX3X and gamma-tubulin.
Dephosphorylated by PP2A-PPP2R5C holoenzyme at Thr-55. SV40 small T antigen inhibits the dephosphorylation by the AC form of PP2A.
May be O-glycosylated in the C-terminal basic region. Studied in EB-1 cell line.
Ubiquitinated by MDM2 and SYVN1, which leads to proteasomal degradation. Ubiquitinated by RFWD3, which works in cooperation with MDM2 and may catalyze the formation of short polyubiquitin chains on p53/TP53 that are not targeted to the proteasome. Ubiquitinated by MKRN1 at Lys-291 and Lys-292, which leads to proteasomal degradation. Deubiquitinated by USP10, leading to its stabilization. Ubiquitinated by TRIM24, RFFL, RNF34 and RNF125, which leads to proteasomal degradation. Ubiquitination by TOPORS induces degradation. Deubiquitination by USP7, leading to stabilization. Isoform 4 is monoubiquitinated in an MDM2-independent manner. Ubiquitinated by COP1, which leads to proteasomal degradation. Ubiquitination and subsequent proteasomal degradation is negatively regulated by CCAR2. Polyubiquitinated by C10orf90/FATS, polyubiquitination is 'Lys-48'-linkage independent and non-proteolytic, leading to TP53 stabilization (By similarity).
Monomethylated at Lys-372 by SETD7, leading to stabilization and increased transcriptional activation. Monomethylated at Lys-370 by SMYD2, leading to decreased DNA-binding activity and subsequent transcriptional regulation activity. Lys-372 monomethylation prevents interaction with SMYD2 and subsequent monomethylation at Lys-370. Dimethylated at Lys-373 by EHMT1 and EHMT2. Monomethylated at Lys-382 by KMT5A, promoting interaction with L3MBTL1 and leading to repress transcriptional activity. Dimethylation at Lys-370 and Lys-382 diminishes p53 ubiquitination, through stabilizing association with the methyl reader PHF20. Demethylation of dimethylated Lys-370 by KDM1A prevents interaction with TP53BP1 and represses TP53-mediated transcriptional activation. Monomethylated at Arg-333 and dimethylated at Arg-335 and Arg-337 by PRMT5; methylation is increased after DNA damage and might possibly affect TP53 target gene specificity.
Sumoylated with SUMO1. Sumoylated at Lys-386 by UBC9.
Cytoplasm. Nucleus. Nucleus>PML body. Endoplasmic reticulum. Mitochondrion matrix. Cytoplasm>Cytoskeleton>Microtubule organizing center>Centrosome.
Note: Interaction with BANP promotes nuclear localization (PubMed:15701641). Recruited into PML bodies together with CHEK2 (PubMed:12810724). Translocates to mitochondria upon oxidative stress (PubMed:22726440). Translocates to mitochondria in response to mitomycin C treatment (PubMed:27323408).
Nucleus. Cytoplasm.
Note: Predominantly nuclear but localizes to the cytoplasm when expressed with isoform 4.
Nucleus. Cytoplasm.
Note: Localized mainly in the nucleus with minor staining in the cytoplasm.
Nucleus. Cytoplasm.
Note: Localized in the nucleus in most cells but found in the cytoplasm in some cells.
Nucleus. Cytoplasm.
Note: Predominantly nuclear but translocates to the cytoplasm following cell stress.
Nucleus. Cytoplasm.
Note: Localized mainly in the nucleus with minor staining in the cytoplasm.
Nucleus. Cytoplasm.
Note: Localized in both nucleus and cytoplasm in most cells. In some cells, forms foci in the nucleus that are different from nucleoli.
Cytoplasm.
Ubiquitous. Isoforms are expressed in a wide range of normal tissues but in a tissue-dependent manner. Isoform 2 is expressed in most normal tissues but is not detected in brain, lung, prostate, muscle, fetal brain, spinal cord and fetal liver. Isoform 3 is expressed in most normal tissues but is not detected in lung, spleen, testis, fetal brain, spinal cord and fetal liver. Isoform 7 is expressed in most normal tissues but is not detected in prostate, uterus, skeletal muscle and breast. Isoform 8 is detected only in colon, bone marrow, testis, fetal brain and intestine. Isoform 9 is expressed in most normal tissues but is not detected in brain, heart, lung, fetal liver, salivary gland, breast or intestine.
The nuclear export signal acts as a transcriptional repression domain. The TADI and TADII motifs (residues 17 to 25 and 48 to 56) correspond both to 9aaTAD motifs which are transactivation domains present in a large number of yeast and animal transcription factors.
Belongs to the p53 family.
研究領域
· Cellular Processes > Cell growth and death > Cell cycle. (View pathway)
· Cellular Processes > Cell growth and death > p53 signaling pathway. (View pathway)
· Cellular Processes > Cell growth and death > Apoptosis. (View pathway)
· Cellular Processes > Cell growth and death > Ferroptosis. (View pathway)
· Cellular Processes > Cell growth and death > Cellular senescence. (View pathway)
· Environmental Information Processing > Signal transduction > MAPK signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Sphingolipid signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Wnt signaling pathway. (View pathway)
· Human Diseases > Drug resistance: Antineoplastic > Endocrine resistance.
· Human Diseases > Drug resistance: Antineoplastic > Platinum drug resistance.
· Human Diseases > Neurodegenerative diseases > Amyotrophic lateral sclerosis (ALS).
· Human Diseases > Neurodegenerative diseases > Huntington's disease.
· Human Diseases > Infectious diseases: Viral > Hepatitis C.
· Human Diseases > Infectious diseases: Viral > Hepatitis B.
· Human Diseases > Infectious diseases: Viral > Measles.
· Human Diseases > Infectious diseases: Viral > Human papillomavirus infection.
· Human Diseases > Infectious diseases: Viral > HTLV-I infection.
· Human Diseases > Infectious diseases: Viral > Herpes simplex infection.
· Human Diseases > Infectious diseases: Viral > Epstein-Barr virus infection.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Overview > Transcriptional misregulation in cancer.
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
· Human Diseases > Cancers: Overview > Proteoglycans in cancer.
· Human Diseases > Cancers: Overview > MicroRNAs in cancer.
· Human Diseases > Cancers: Specific types > Colorectal cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Pancreatic cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Endometrial cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Glioma. (View pathway)
· Human Diseases > Cancers: Specific types > Prostate cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Thyroid cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Basal cell carcinoma. (View pathway)
· Human Diseases > Cancers: Specific types > Melanoma. (View pathway)
· Human Diseases > Cancers: Specific types > Bladder cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Chronic myeloid leukemia. (View pathway)
· Human Diseases > Cancers: Specific types > Small cell lung cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Non-small cell lung cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Breast cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Hepatocellular carcinoma. (View pathway)
· Human Diseases > Cancers: Specific types > Gastric cancer. (View pathway)
· Human Diseases > Cancers: Overview > Central carbon metabolism in cancer. (View pathway)
· Organismal Systems > Aging > Longevity regulating pathway. (View pathway)
· Organismal Systems > Nervous system > Neurotrophin signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Thyroid hormone signaling pathway. (View pathway)
参考文献
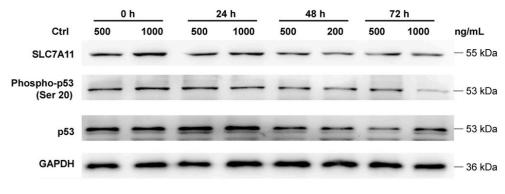
Application: WB Species: Human Sample: U251 cells
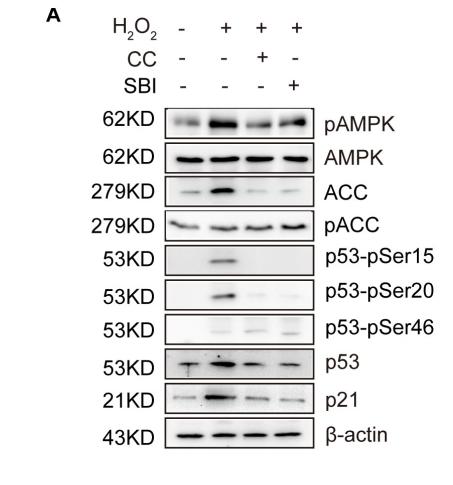
Application: WB Species: Human Sample: VSMCs
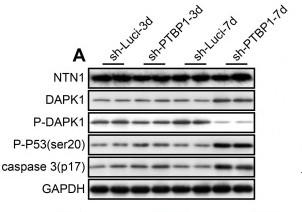
Application: WB Species: mouse Sample: zygotes
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.