PKM2 Antibody - #AF5234
製品説明
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
引用形式: Affinity Biosciences Cat# AF5234, RRID:AB_2837720.
折りたたみ/展開
CTHBP; Cytosolic thyroid hormone binding protein; Cytosolic thyroid hormone-binding protein; KPYM_HUMAN; MGC3932; OIP 3; OIP-3; OIP3; OPA interacting protein 3; Opa-interacting protein 3; p58; PK muscle type; PK, muscle type; PK2; PK3; PKM; PKM2; pykm; Pyruvate kinase 2/3; Pyruvate kinase 3; Pyruvate kinase isozymes M1/M2; Pyruvate kinase muscle; Pyruvate kinase muscle isozyme; pyruvate kinase PKM; Pyruvate kinase, muscle 2; TCB; THBP1; Thyroid hormone binding protein 1; Thyroid hormone binding protein cytosolic; Thyroid hormone-binding protein 1; Tumor M2 PK; Tumor M2-PK;
免疫原
A synthesized peptide derived from human PKM2, corresponding to a region within the internal amino acids.
Specifically expressed in proliferating cells, such as embryonic stem cells, embryonic carcinoma cells, as well as cancer cells.
- P14618 KPYM_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSKPHSEAGTAFIQTQQLHAAMADTFLEHMCRLDIDSPPITARNTGIICTIGPASRSVETLKEMIKSGMNVARLNFSHGTHEYHAETIKNVRTATESFASDPILYRPVAVALDTKGPEIRTGLIKGSGTAEVELKKGATLKITLDNAYMEKCDENILWLDYKNICKVVEVGSKIYVDDGLISLQVKQKGADFLVTEVENGGSLGSKKGVNLPGAAVDLPAVSEKDIQDLKFGVEQDVDMVFASFIRKASDVHEVRKVLGEKGKNIKIISKIENHEGVRRFDEILEASDGIMVARGDLGIEIPAEKVFLAQKMMIGRCNRAGKPVICATQMLESMIKKPRPTRAEGSDVANAVLDGADCIMLSGETAKGDYPLEAVRMQHLIAREAEAAIYHLQLFEELRRLAPITSDPTEATAVGAVEASFKCCSGAIIVLTKSGRSAHQVARYRPRAPIIAVTRNPQTARQAHLYRGIFPVLCKDPVQEAWAEDVDLRVNFAMNVGKARGFFKKGDVVIVLTGWRPGSGFTNTMRVVPVP
種類予測
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
研究背景
Glycolytic enzyme that catalyzes the transfer of a phosphoryl group from phosphoenolpyruvate (PEP) to ADP, generating ATP. Stimulates POU5F1-mediated transcriptional activation. Plays a general role in caspase independent cell death of tumor cells. The ratio between the highly active tetrameric form and nearly inactive dimeric form determines whether glucose carbons are channeled to biosynthetic processes or used for glycolytic ATP production. The transition between the 2 forms contributes to the control of glycolysis and is important for tumor cell proliferation and survival. Promotes in a STAT1-dependent manner, the expression of the immune checkpoint protein CD274 in ARNTL/BMAL1-deficient macrophages (By similarity).
ISGylated.
Under hypoxia, hydroxylated by EGLN3.
Acetylation at Lys-305 is stimulated by high glucose concentration, it decreases enzyme activity and promotes its lysosomal-dependent degradation via chaperone-mediated autophagy.
FGFR1-dependent tyrosine phosphorylation is reduced by interaction with TRIM35.
Cytoplasm. Nucleus.
Note: Translocates to the nucleus in response to different apoptotic stimuli. Nuclear translocation is sufficient to induce cell death that is caspase independent, isoform-specific and independent of its enzymatic activity.
Specifically expressed in proliferating cells, such as embryonic stem cells, embryonic carcinoma cells, as well as cancer cells.
Belongs to the pyruvate kinase family.
研究領域
· Human Diseases > Endocrine and metabolic diseases > Type II diabetes mellitus.
· Human Diseases > Infectious diseases: Viral > Human papillomavirus infection.
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
· Human Diseases > Cancers: Overview > Central carbon metabolism in cancer. (View pathway)
· Metabolism > Carbohydrate metabolism > Glycolysis / Gluconeogenesis.
· Metabolism > Nucleotide metabolism > Purine metabolism.
· Metabolism > Carbohydrate metabolism > Pyruvate metabolism.
· Metabolism > Global and overview maps > Metabolic pathways.
· Metabolism > Global and overview maps > Carbon metabolism.
· Metabolism > Global and overview maps > Biosynthesis of amino acids.
· Organismal Systems > Endocrine system > Glucagon signaling pathway.
参考文献
Application: WB Species: human Sample: MIA PaCa-2 cells
Application: WB Species: Human Sample: pancreatic cancer tissue
Application: WB Species: Human Sample: MCF-7 cells
Application: WB Species: Mouse Sample: BMSCs
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.






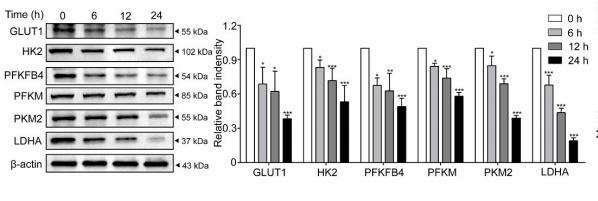
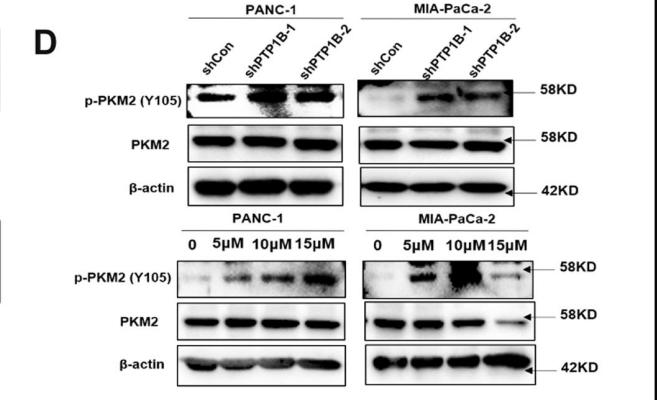
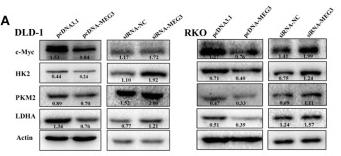
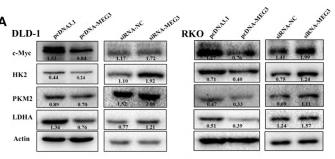


![Figure 5 BHD inhibited protein and mRNA expression of pyruvate kinase M2 (PKM2) and hypoxia-inducible factor-1 alpha (HIF-1α) as well as its target genes (GLUT1, PDK1, lactate dehydrogenase A [LDHA]) in aortas after high fat diet treatment. (a) Protein Levels of PKM2 were measured by Western blot analysis, and quantitative analysis was performed on the corresponding bands (n = 8–9). (b) Protein Levels of HIF-1α was measured by WB, and quantitative analysis was performed on the corresponding bands (n = 9). (c) The mRNA levels of GLUTA, (d) PDK1, and (e) LDHA in aortas (n = 6) were measured by qRT-PCR. Means ± standard error of mean. *P < 0.05, **P < 0.01, ***P < 0.001 versus Con; #P < 0.05, ##P < 0.01 versus Mod. PKM2 Antibody - Figure 5 BHD inhibited protein and mRNA expression of pyruvate kinase M2 (PKM2) and hypoxia-inducible factor-1 alpha (HIF-1α) as well as its target genes (GLUT1, PDK1, lactate dehydrogenase A [LDHA]) in aortas after high fat diet treatment.](http://img.affbiotech.cn/uploads/202409/5ce509e907ddfd09b9be49206226cfbb.png)

